# Copyright 2016 Nicolas P. Rougier - MIT License
# https://github.com/rougier/matplotlib-tutorial
fig = plt.figure(figsize=(10, 7))
ax = fig.add_axes([0.2, 0.17, 0.68, 0.7], aspect=1)
# Main data
X = np.linspace(0.5, 3.5, 100)
Y1 = 3 + np.cos(X)
Y2 = 1 + np.cos(1 + X / 0.75) / 2
Y3 = np.random.uniform(Y1, Y2, len(X))
ax.fill_between(X, Y1, Y2, color="C0", alpha=0.25)
ax.plot(X, Y1, c="C0", label="Blue signal", linewidth=2)
ax.plot(X, Y2, c="C0", linewidth=2)
ax.plot(X[::3], Y3[::3], linewidth=0, marker='o', markerfacecolor="w",
markeredgecolor="C1", markeredgewidth=1.5, label="Markers")
# Setup axes
ax.set_xlim(0, 4); ax.set_ylim(0, 4.5)
ax.tick_params(which="major", width=1.5, length=6)
ax.tick_params(which="minor", width=1.0, length=3)
ax.xaxis.set_major_locator(plt.MultipleLocator(1.0))
ax.xaxis.set_minor_locator(plt.MultipleLocator(0.25))
ax.yaxis.set_major_locator(plt.MultipleLocator(1.0))
ax.yaxis.set_minor_locator(plt.MultipleLocator(0.25))
ax.grid(True, linestyle='--', alpha=0.3)
ax.set_xlabel("X axis label", fontsize=14)
ax.set_ylabel("Y axis label", fontsize=14)
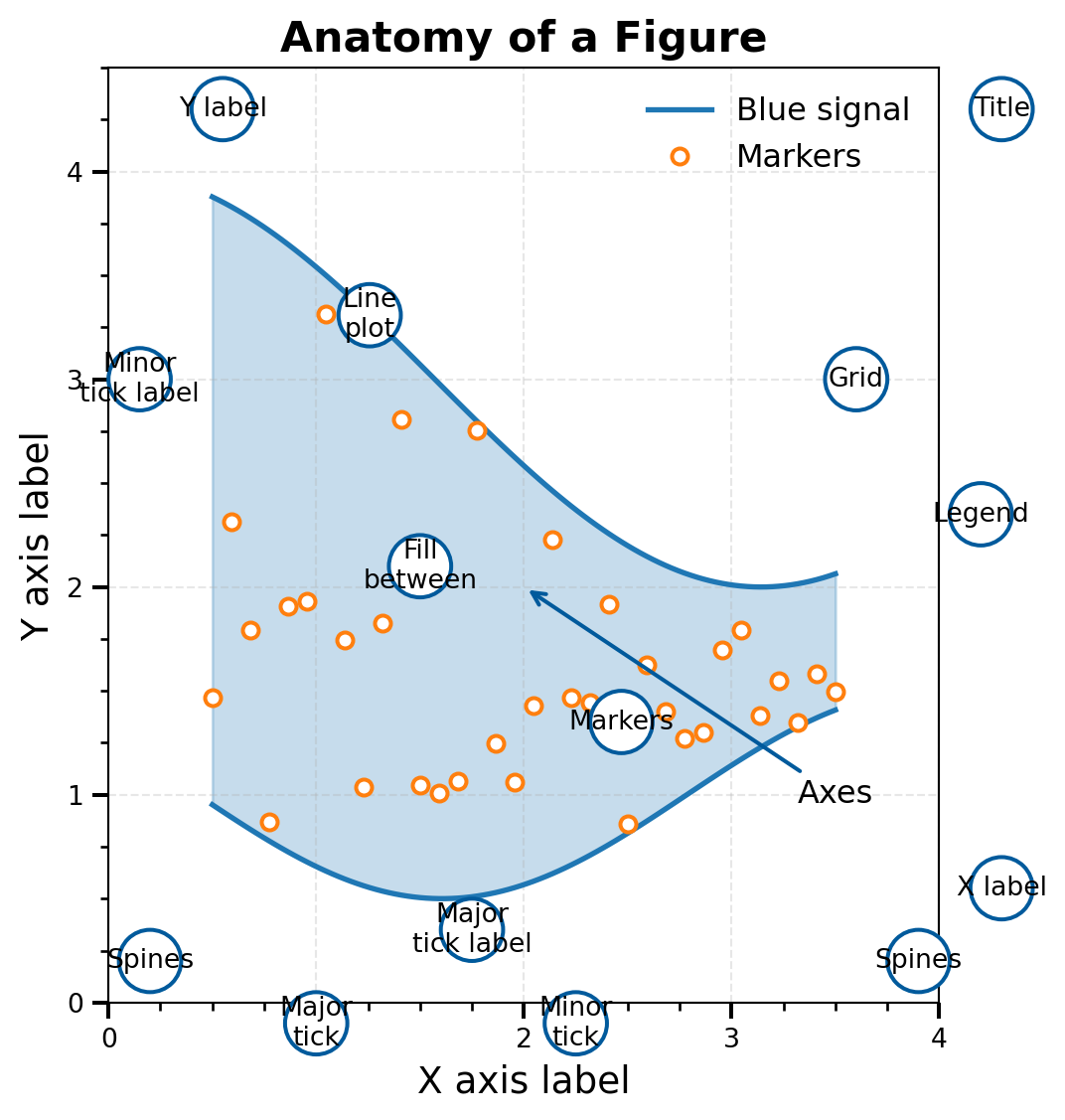
ax.set_title("Anatomy of a Figure", fontsize=16, fontweight="bold")
ax.legend(loc="upper right", frameon=False, fontsize=12)
# Annotation helper
def circle(x, y, radius=0.15):
from matplotlib.patches import Circle as C
c = C((x, y), radius, clip_on=False, zorder=10, lw=1.5,
ec="#005A9C", fc="white")
ax.add_artist(c)
def text(x, y, t):
ax.text(x, y, t, ha="center", va="center", size=10,
zorder=20, family="Source Sans Pro")
# All annotations
circle(X[25], Y1[25]); text(X[25], Y1[25], "Line\nplot")
circle(X[65], Y3[65]); text(X[65], Y3[65], "Markers")
circle(1.5, 2.1); text(1.5, 2.1, "Fill\nbetween")
circle(4.3, 4.3); text(4.3, 4.3, "Title")
circle(4.3, 0.55); text(4.3, 0.55, "X label")
circle(0.55, 4.3); text(0.55, 4.3, "Y label")
circle(4.2, 2.35); text(4.2, 2.35, "Legend")
circle(1.75, 0.35); text(1.75, 0.35, "Major\ntick label")
circle(2.25, -0.1); text(2.25, -0.1, "Minor\ntick")
circle(1.0, -0.1); text(1.0, -0.1, "Major\ntick")
circle(0.15, 3.0); text(0.15, 3.0, "Minor\ntick label")
circle(3.6, 3.0); text(3.6, 3.0, "Grid")
circle(3.9, 0.2); text(3.9, 0.2, "Spines")
circle(0.2, 0.2); text(0.2, 0.2, "Spines")
ax.annotate("Axes", xy=(2.0, 2.0), xycoords="data", xytext=(3.5, 1.0),
fontsize=12, ha="center", va="center",
arrowprops=dict(arrowstyle="->", color="#005A9C", lw=1.5))
ax.annotate("Figure", xy=(4.85, 4.7), xycoords="data", xytext=(4.1, 4.7),
fontsize=12, ha="left", va="center",
arrowprops=dict(arrowstyle="->", color="#005A9C", lw=1.5))
plt.show()